We explain the basic idea of probabilistic machine learning: data are drawn from a distribution, and machine learning learns this distribution. Notation
Define sample space as the set of all possible outcomes, denoted as $\Omega$. Define the measure $(\Omega, \mathcal{F}, P)$ A sample is a single element $\omega \in \Omega$ An event is a subset $A \subseteq \Omega$ (more precisely, $A \in \mathcal{F}$) A random variable $X: \Omega \rightarrow \mathbb{R}^d$ is a measurable function. A value $x \in \mathbb{R}^d$ is a specific point in the target space. With $x$, an event is a measurable set, denoted
\[{X=x}:={\omega \in \Omega: X(\omega)=x}\] which is the pre-image $X^{-1}({x})$ A probability measure is a triple $(\Omega, \mathcal{F}, P)$ where $P: \mathcal{F} \rightarrow[0,1]$ A probability of event is $P(X=x)\in [0,1]$ A distribution of $X$, denoted as $P_X$, is a pushforward measure induced by a random variable: When we have a random variable $X: \Omega \rightarrow \mathbb{R}^d$, it induces a probability measure on $\mathbb{R}^d$ :
\[P_X(A):=P\left(X^{-1}(A)\right)=P({\omega \in \Omega: X(\omega) \in A})\] for measurable sets $A \subseteq \mathbb{R}^d$.
We have a problem: an event can happen even if its probability is 0.
For example, in continuous $X$:
\[P(X=x)=P({\omega \in \Omega: X(\omega)=x})=0\] So we can’t assign positive probability to individual points. To solve this, we introduce the Probability Density Function (PDF) The PDF $p_X(x)$ is a function such that:
\[P(X \in A)=\int_A p_X(x) d x\] for any measurable set $A$.
Note:
- $p_X(x)$ is not a probability. It’s a density, so it can be greater than 1
- $p_X(x) d x$ represents the probability of being in an infinitesimal region around $x$. Formally
\[p_X(x)=\lim_{\epsilon \rightarrow 0} \frac{P(x \leq X \leq x+\epsilon)}{\epsilon}\] Their relation:
\[P(Y=y)=P_Y({y})=\int_{\lbrace y\rbrace} p_Y(y) d y=0\] In this essay, we denote a latent variable as $X$, observation data as $Y$.
$p(\cdot)$
This is one of the most abused notations in probability and machine learning. Strictly speaking, the lowercase $p$ denotes PDF, the capital $P$ denotes probability, the lowercase $x$ denotes a value of random variable, the capital $X$ means random variable.
| Notation | Actual Meaning | Function Type |
| $p(X)$ | $p_X(x)$ | Marginal density of X |
| $p(Y)$ | $p_Y(y)$ | Marginal density of Y |
| $p(X, Y)$ | $p_{X,Y}(x, y)$ | Joint density |
| $p(Y \mid X)$ | $p_{Y\mid X}(y \mid x)$ | Conditional density |
For Bayes’ theorem (more rigorous explanation check Probability-1): In statistics, usually we write $p(X, Y)=p(Y \mid X) p(X)$, means “the relationship between these density functions” In ML, usually we write $p(x, y)=p(y \mid x) p(x)$, is shorthand for
\[p_{X, Y}(x, y)=p_{Y \mid X}(y \mid x) \cdot p_X(x)\] where:
- $p_{X, Y}: \mathbb{R}^{d_1} \times \mathbb{R}^{d_2} \rightarrow[0, \infty)$ is joint density function
- $p_{Y \mid X}: \mathbb{R}^{d_2} \times \mathbb{R}^{d_1} \rightarrow[0, \infty)$ is conditional density function
- $p_X: \mathbb{R}^{d_1} \rightarrow[0, \infty)$ is marginal density function
- $x, y$ are specific values these random variables can take.
If condition is an observed $x$, we usually write $p(x, Y)=p(Y \mid x) p(x)$
But in ML, lowercase $p$ and $x$ are used for everything: $p(x)$ can mean density function, or distribution, or probability masses $p(X=x)$. $p(X)$ can denote distribution, or density function. The meaning depends on context.
I can understand what $Y$ means, it’s our observation, the data we collected, but what do you mean by “ latent variable? Is it what we observe or what we believe?
Here is the abstract part of probability theory: we don’t care (and usually don’t know) what $X$ really is, we can think of it as an object that satisfies certain properties, for example:
- $X\sim \mathcal{N}(0,I)$
- $x\in X$, $P(x)=0.1$
The Learning Story: take VAE for example
What’s our observation (What do we know)?
We observe dataset $Y=\lbrace y_1, y_2, \ldots, y_n\rbrace$
What’s our assumption?
We assume the distribution of the latent variable $X$, for example, $p(X)=\mathcal{N}(0, I)$, called prior. Also we assume $Y \mid X$ follows some distribution, for example $p(Y \mid X=x)= \mathcal{N}\left(f(x), \sigma^2 I\right)$, called likelihood.
Why we need to specify $X=x$ ?
Because our assumption is a causal/generative structure where $Y$ depends on $X$. The distribution of $Y$ changes based on what value $X$ takes.
What do we want to know?
We want to learn the parameters of function $f$ in $\mathcal{N}\left(f(x), \sigma^2 I\right)$.
Examples of $f$:
Linear model: $f(x)=W x+b$
- Learn: weight matrix $W$ and bias $b$
- This is Probabilistic PCA or Factor Analysis
Neural network: $f(x)=\mathrm{NN}_\theta(x)$
- Learn: neural network weights $\theta$
- This is a Variational Autoencoder (VAE)
Gaussian Process: $f \sim \mathcal{G} \mathcal{P}(m, k)$
- Learn: kernel parameters
- This is Gaussian Process Latent Variable Model (GPLVM)
Set up
Our question: What’s the distribution of $Y$? Or, what’s the probability of observing $y\in Y$, i.e., $p(y)$?
The idea: $Y$ is generated by a latent variable $X$. Let’s write $p(y)$ using $X$. In a more concrete (or abstract?) saying: let’s detect $Y$ using $X$ in a probabilistic way.
Prior (assumed, fixed):
\[p(X)=\mathcal{N}(0, I)\] Likelihood (form assumed, parameters learned):
\[p(Y \mid X=x)=\mathcal{N}\left(f_\theta(x), \sigma^2 I\right)\] where $f_\theta$ is a neural network with weights $\theta$ Observe:
\[Y = \lbrace y_1, y_2, \ldots, y_n\rbrace\] Unknown (to be learned):
\[\theta=\lbrace W_1, b_1, W_2, b_2, \ldots\rbrace\]
REMARK: Technically all $p(\cdot)$ and $q(\cdot)$ in formulas are density not probability, but densities satisfy the same algebraic rules as probabilities (Bayes, marginalization, chain rule), so we say they are probability, but calculate them as density. Why density? We will see it later…
By Bayes’ theorem, the joint distribution:
\[p_\theta(x, y)=p_\theta(y \mid x) \cdot p(x)\] where:
- $p(x)=\mathcal{N}(0, I)$ is our prior
- $p_\theta(y \mid x)=\mathcal{N}\left(f_\theta(x), \sigma^2 I\right)$ is our likelihood, parameterized by $\theta$
The marginal (called marginal likelihood, or evidence) is a integral:
\[p_\theta(y)=\int p_\theta(y \mid x) \cdot p(x) d x\] Back to the question: What’s the probability of observing $y$ under parameter $\theta$? The answer is: We follow the trace $x$ generates $y$. But we need to consider all possible latent $x$ that could have generated $y$.
Objective: Maximum Likelihood Estimation (MLE) of the parameters $\theta$. Let’s formalize this objective.
Define the Likelihood Function:
\[\mathcal{L}(\theta ; y):=p_\theta(y)\] A bit confusing: $\mathcal{L}(\theta ; y)=p_\theta(y)$ has different interpretations:
- As a function of $y$ with $\theta$ fixed : It’s a probability distribution
- As a function of $\theta$ with $y$ fixed : It’s a likelihood function In learning story $y$ is known, we want to learn $\theta$, so it is a likelihood function. Likelihood is NOT a probability distribution over $\theta$, its integral generally is not 1, it’s just a positive function of $\theta$.
Maybe more confusing, we have different types of likelihood:
- Conditional likelihood: $p_\theta(y \mid x)$ : probability of $y$ given $x$ and $\theta$
- Marginal likelihood: $p_\theta(y)=\int p_\theta(y \mid x) p(x) d x$: probability of $y$ given $\theta$ (marginalizing $x$ )
- Likelihood function: $\mathcal{L}(\theta ; y)=p_\theta(y)$ : same as (2), but viewed as a function of $\theta$
Recall our Objective: Maximum Likelihood Estimation (MLE), find $\theta$ that maximizes the likelihood of the observed data.
Which likelihood?
First we compare MLE and MAP
MLE (Maximum Likelihood Estimation) - Frequentists’ perspective Objective: Find $\theta$ that maximizes likelihood
\[\hat{\theta}_{M L E}=\arg \max _\theta p(y \mid \theta)\] Philosophy:
- $\theta$ is a fixed (unknown, we need to learn to step by step) value
- No probability distribution over $\theta$
- Find single “best” estimate
MAP (Maximum A Posteriori) - Bayesians’ perspective Objective: Find $\theta$ that maximizes posterior
\[\hat{\theta}_{M A P}=\arg \max _\theta p(\theta \mid y)\] Philosophy:
- $\theta$ is a random variable with distribution
- Prior beliefs about $\theta$ encoded in $p(\theta)$
- Update beliefs using data
DIGRESSION: The distinction between frequentist and Bayesian perspectives are also reflected in the symbols. There are two ways to denote conditional probability: $p_\theta(y)$ and $p(y \mid \theta)$, numerically they’re the same, but differ in philosophy. $p_\theta(y)$ : Frequentist view:
\[p(\theta \mid y)=\frac{p(y \mid \theta) p(\theta)}{p(y)}\] Modern machine learning commonly use hybrid approach, which is also called Type II Maximum Likelihood: Maximize $p_\theta(y)=\int p_\theta(y \mid x) p(x) d x$ over $\theta$
Bayes’ Equation: Inferring Latents Given Data
\[p_\theta(x \mid y)=\frac{p_\theta(y \mid x) \cdot p(x)}{p_\theta(y)}\] where
- Posterior: $p_\theta(x \mid y)$ - what we want (latent given data)
- Likelihood: $p_\theta(y \mid x)$ - conditional likelihood
- Prior: $p(x)$ - prior on latents
- Evidence: $p_\theta(y)$ - marginal likelihood
The likelihood Function: $\mathcal{L}(\theta ; y)=p_\theta(y)$, with different perspectives of $y$ and $\theta$ , is
- Probability of data y given parameters $\theta$
- Notation: $p(y \mid \theta)$ or $p_\theta(y)$
- A function of $\theta$ (with $y$ fixed)
- NOT a probability distribution over $\theta$
So, the “likelihood” in Maximum Likelihood Estimation, is NOT the (conditional) likelihood is Bayes’s equation related to $x$ and $y$, instead, it is the evidence, aka marginal likelihood. Maximum Likelihood Estimation actually maximize the likelihood function $\mathcal{L}(\theta ; y)$ , which is $p_\theta(y)$ viewed as a function of $\theta$.
Collect all the observations $y_i$, our objective is:
\[\max _\theta \prod_{i=1}^n p_\theta\left(y_i\right)=\max _\theta \prod_{i=1}^n \int p_\theta\left(y_i \mid x\right) p(x) d x\] Or in log form:
\[\max _\theta \sum_{i=1}^n \log p_\theta\left(y_i\right)=\max _\theta \sum_{i=1}^n \log \int p_\theta\left(y_i \mid x\right) p(x) d x\] But in general, this integral is intractable.
Why?
$p_\theta\left(y_i \mid x\right)$ contains a neural network; basically, we can believe that everything that contains a neural network is not analytically integrable. Also, we can’t use the sampling method (Monte Carlo), if we want to sample from the prior $p(x)$ to approximate
\[\log \int p_\theta(y \mid x) p(x) d x \approx \log \frac{1}{L} \sum_{l=1}^L p_\theta\left(y \mid x^{(l)}\right)\] where $x^{(l)} \sim p(x)=\mathcal{N}(0, I)$. We want $p_\theta\left(y \mid x^{(l)}\right)$ to be as high as possible, so it’s an effective sample. But the latent space (all possible $x$ ) is very broad. The set of $x$ that generate a specific $y$ (for example, an image )is very narrow. Most random samples from $p(x)$ give tiny $p_\theta\left(y \mid x^{(l)}\right)$, making the sampling method exponentially inefficient.
One guiding principle from engineering: we don’t need to be the best, we just need to be good enough.
Evidence Lower Bound (ELBO)
We make another assumption: $q_\phi(x \mid y)$ as a known distribution. For example $q_\phi(x \mid y)=\mathcal{N}\left(\mu_\phi(y), \operatorname{diag}\left(\sigma_\phi^2(y)\right)\right)$ , its only parameter is $\phi$.
The variational trick: Using $q_\phi(x \mid y)$ to approximate the true posterior $p_\theta(x \mid y)$ .
Multiply and divide by $q_\phi(x \mid y)$ inside the integral:
\[\begin{aligned} \log p_\theta(y) &=\log \int p_\theta(y, x) d x\\ & =\log \int q_\phi(x \mid y) \frac{p_\theta(y, x)}{q_\phi(x \mid y)} d x \\ & =\log \mathbb{E}_{q_\phi(x \mid y)}\left[\frac{p_\theta(y, x)}{q_\phi(x \mid y)}\right]\\ &\geq \mathbb{E}_{q_\phi(x \mid y)}\left[\log \frac{p_\theta(y, x)}{q_\phi(x \mid y)}\right] \quad \text{(by Jensen's inequality, since } \log \text{ is concave)}\\ &=\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y, x)\right]-\mathbb{E}_{q_\phi(x \mid y)}\left[\log q_\phi(x \mid y)\right]\\ &=\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)+\log p(x)\right]-\mathbb{E}_{q_\phi(x \mid y)}\left[\log q_\phi(x \mid y)\right]\\ &=\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]+\mathbb{E}_{q_\phi(x \mid y)}\left[\log p(x)-\log q_\phi(x \mid y)\right]\\ &=\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]-D_{K L}\left(q_\phi(x \mid y) \lVert p(x)\right) \end{aligned}\] where $\mathbb{E}{q\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]-D_{K L}\left(q_\phi(x \mid y) \lVert p(x)\right)$ is called ELBO.
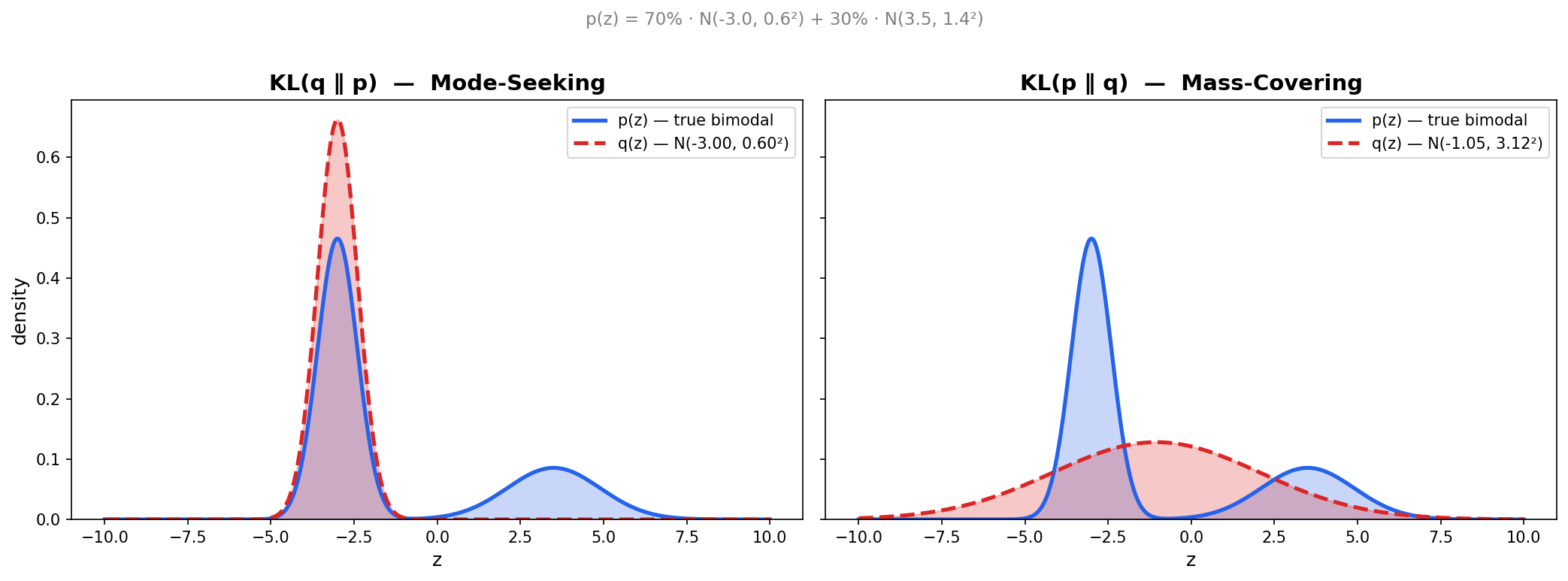
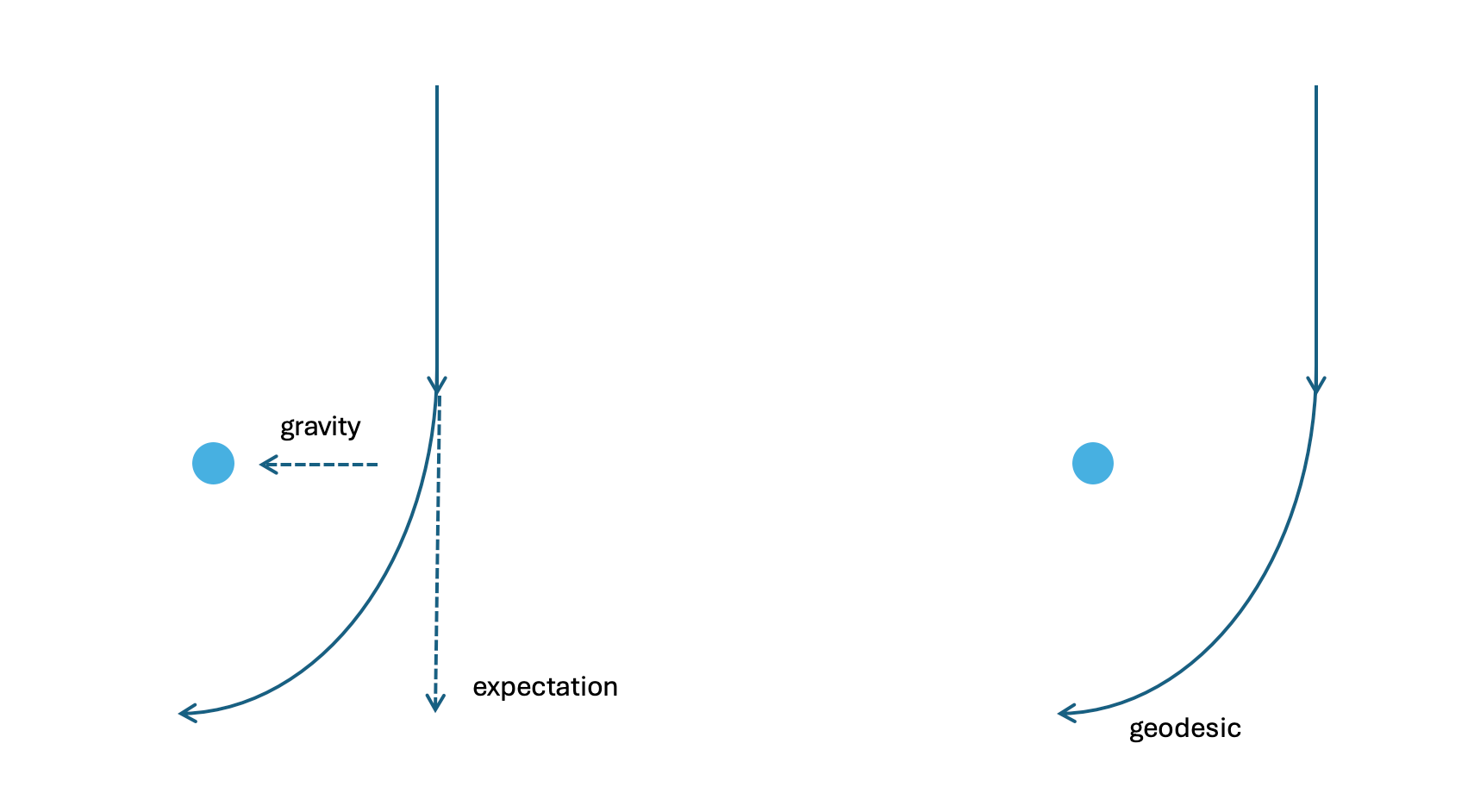
Note: Here we define KL divergence $D_{KL}$, which measures the difference between two probability distributions (but not a distance). There are many ways to measure differences between probabilities, KL divergence is a good one: analogously, it is the “Euclidean distance” in probability space.
REMARK: There is another sloppy notation. Technically $p(\cdot)$ and $q(\cdot)$ are density function, not probability. When we write
\[\int q_\phi(x \mid y) \cdot g(x) d x=\mathbb{E}_{q_\phi(x \mid y)}[g(X)]\] what we actually mean is
\[\int q_\phi(x \mid y) \cdot g(x) d x=\mathbb{E}_{X \sim Q_\phi}[g(X)]\] where $Q_\phi$ is the probability measure, the $q_\phi(x \mid y)$ is its density, such that $Q_\phi(d x)=q_\phi(x \mid y) d x$ Most time we can interchangeably use probability and density. But in some cases, for example in this expectation trick, $p(\cdot)$ must be density. Another non-trivial case is reparameterization: If $Y=h(X)$ for some transformation $h$ , then
- Probability is preserved: $P(Y \in h(A))=P(X \in A)$
- Density transforms with a Jacobian:
\[p_Y(y)=p_X\left(h^{-1}(y)\right) \cdot\lvert\frac{d h^{-1}}{d y}\rvert\] Densities are not invariant under reparameterization.
Now we know $\log p_\theta(y)\geq \mathrm{ELBO}$, so ELBO is a lower bound on $\log p_\theta(y)$,
Next we recover $\log p_\theta(y)$ from ELBO
\[\mathrm{ELBO}=\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)-\log q_\phi(x \mid y)+\log p(x)\right]\] Substitute using Bayes’ equation:
\[\mathrm{ELBO}=\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y)+\log p_\theta(x \mid y)-\log q_\phi(x \mid y)\right]\] Since $\log p_\theta(y)$ doesn’t depend on $x$, it’s constant with respect to the expectation over $q_\phi(x \mid y)$ :
\[\mathrm{ELBO}=\log p_\theta(y)+\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(x \mid y)-\log q_\phi(x \mid y)\right]\] This gives us the key equation:
\[\log p_\theta(y)=\operatorname{ELBO}(\theta, \phi ; y)+D_{K L}\left(q_\phi(x \mid y) \lVert p_\theta(x\mid y)\right)\] where ELBO is a scalar-valued function of parameters $\theta$ and $\phi$.
Since $D_{K L} \geq 0$, the same conclusion as before: ELBO is always a lower bound on $\log p_\theta(y)$. When $q_\phi(x \mid y)=p_\theta(x \mid y)$, i.e., our approximate posterior exactly matches the true posterior, ELBO equals the true log-likelihood.
We observe an interesting phenomena in this equation. Since the left side is fixed when we vary $\phi$ :
\[\frac{\partial}{\partial \phi} \log p_\theta(y)=0\] Therefore:
\[\frac{\partial}{\partial \phi} \operatorname{ELBO}(\theta, \phi ; y)=-\frac{\partial}{\partial \phi} D_{K L}\left(q_\phi(x \mid y) \lVert p_\theta(x \mid y)\right)\] When we increase ELBO by changing $\phi$ , we decreases KL divergence simultaneously!
But all components of this equation depend on $\theta$. When we increase ELBO by changing $\theta$, $\log p_\theta(y)$ increases, but at the same time, the true posterior $p_\theta(x \mid y)$ is moving, so KL might increase or decrease. But overall, we’re making the model better!
Now, our objective becomes
\[\max _{\theta, \phi} \operatorname{ELBO}(\theta, \phi ; y)\] Methodology
Recall the ELBO formula:
\[\operatorname{ELBO}(\theta, \phi ; y)=\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]-D_{K L}\left(q_\phi(x \mid y) \lVert p(x)\right)\] This looks more complex, we need to optimize with respect to both $\theta$ and $\phi$. Is it tractable?
Yes! We untangle it item by item.
KL item
The KL part is relatively easy. For example if we take both Gaussian:
- $q_\phi(x \mid y)=\mathcal{N}\left(\mu_\phi(y), \operatorname{diag}(\sigma_\phi^2(y))\right)$
- $p(x)=\mathcal{N}(0, I)$ The KL divergence (for diagonal covariance Gaussian vs standard normal) is:
\[D_{K L}=\frac{1}{2} \sum_{j=1}^d\left(\mu_j^2+\sigma_j^2-\log \sigma_j^2-1\right)\] where $\mu=\mu_\phi(y)$ and $\sigma^2=\sigma_\phi^2(y)$. Note: this closed form assumes $q_\phi$ has diagonal covariance; the general case with a full covariance matrix is more complex. This is differentiable with respect to $\phi!$ We can compute gradients directly.
Expectation item
The expectation item needs to be more careful. By definition
\[\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]=\int q_\phi(x \mid y)\log p_\theta(y \mid x)dx\] A little break…
Let’s clarify notations before we go further.
What is $q_\phi(x \mid y) ?$
It’s our assumption used to surrogate the true posterior $p_\theta(x \mid y)$ where
- $\phi$ is the parameter that defines the function
- $y$ is the observation, aka the conditional variable, which is given/fixed when we write this expression
- $x$ is the argument of the function. This is what the density function takes as input and produces density values for, $x$ ranges over all possible latent codes We can think of it as a function:
\[q_\phi(x \mid y): \mathbb{R}^{d_{\text {latent }}} \rightarrow[0, \infty)\] Given fixed $\phi$ (parameters) and fixed $y$ (observation), this is a function of $x$ :
- Input: a latent code $x$
- Output: density value $q_\phi(x \mid y)$
What is $\log p_\theta(y \mid x)$?
Similarly we can think it as a Function:
\[p_\theta(y \mid x): \mathbb{R}^{d_{\text {obs }}} \rightarrow[0, \infty)\] Given fixed $\theta$ and fixed $x$, this is a function of $y$ :
- Input: an observation $y$
- Output: density value $p_\theta(y \mid x)$
What is $\int q_\phi(x \mid y)\log p_\theta(y \mid x)dx$ ?
It is the average value of $\log p_\theta(y \mid x)$ when $x$ is distributed according to $q_\phi(x \mid y)$ . We’re not picking a specific $x$ . We’re computing a weighted average over all possible $x$ values, where the weights are given by $q_\phi(x \mid y)$.
Expectation item again
By definition
\[\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]=\int q_\phi(x \mid y)\log p_\theta(y \mid x)dx\] Parameter $\theta$ is relatively easy: $\nabla_\theta \mathbb{E}{q\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]$. We focus on $\phi$.
First we try to find what is the best $\phi$ for this item , or equally , update $\phi$ via the trajectory
\[\begin{gathered} \nabla_\phi\mathbb{E}_{q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]\\ =\nabla_\phi \int q_\phi(x \mid y) \log p_\theta(y \mid x) d x \end{gathered}\] Recall $p_\theta(y \mid x)=\mathcal{N}\left(f_\theta(x), \sigma^2 I\right)$ , since $f_\theta(x)$ doesn’t depend on $\phi$,
\[=\int \nabla_\phi q_\phi(x \mid y) \cdot \log p_\theta(y \mid x) d x\] A common choice $q_\phi(x \mid y)=\mathcal{N}\left(\mu_\phi(y), \sigma_\phi^2(y)\right)$, but this is still hard because $\mu$ and $\sigma$ are neural networks (encoder). The main issue is that we are integrating target variable.
How can we avoid integration?
We hope the integration can be written as expectation, then expectation can be approximated by Monte Carlo.
Reparameterization trick
This is a really smart and elegant idea.
What can we usually do with a distribution? We sample a point from it. But this “sample” operation is really confusing—how do we do it? Ideally, the sampled element should preserve all information about its distribution, so we know this point is truly sampled from the correct distribution, not randomly generated. Another issue is that, in machine learning, we need to update the parameters of the distribution using this sampled point. How can we achieve that?
The problem: $\phi$ appears in the distribution we’re sampling from ($q_\phi$), so we can’t differentiate through the sampling.
The solution: reparameterize so that $\phi$ appears only in a deterministic transformation of a fixed noise distribution.
In mathematics, we constantly use different perspectives to view identical objects. For example, $q_\phi(x \mid y)$ is both a function of $x$ with codomain $[0,\infty)$, and also a distribution $\mathcal{N}\left(\mu_\phi(y), \sigma_\phi^2(y)\right)$ with parameter $\phi$. The value of the function $q_\phi$ gives $x$, is identical to, the probability of sampling $x$ from the distribution $q_\phi$.
Based on this idea, instead of sampling:
\[x \sim q_\phi(x \mid y)=\mathcal{N}\left(\mu_\phi(y), \sigma_\phi^2(y)\right)\] We use a deterministic function:
\[x=g(\phi, \epsilon, y)=\mu_\phi(y)+\sigma_\phi(y) \cdot \epsilon\] where $\epsilon \sim p(\epsilon)=\mathcal{N}(0, I)$ is a fixed noise variable that doesn’t depend on $\phi$. This is valid because they give the same distribution for $X$. Instead of sampling $x$, we now sample $\epsilon$.
The original expectation:
\[\mathbb{E}_{x \sim q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]=\int q_\phi(x \mid y) \log p_\theta(y \mid x) d x\] Can be rewritten as:
\[\mathbb{E}_{\epsilon \sim p(\epsilon)}\left[\log p_\theta(y \mid g(\phi, \epsilon, y))\right]=\int p(\epsilon) \log p_\theta(y \mid g(\phi, \epsilon, y)) d \epsilon\] We changed variables in the integral!
Key difference: Now $\phi$ appears only in the integrand $g(\phi, \epsilon, y)$, NOT in the distribution $p(\epsilon)$ . Since $p(\epsilon)$ doesn’t depend on $\phi$ , we can rewrite gradient of expectation as the expectation of gradient
\[\begin{gathered} \nabla_\phi\mathbb{E}_{\epsilon \sim p(\epsilon)}\left[\log p_\theta(y \mid g(\phi, \epsilon, y))\right] \\ =\mathbb{E}_{\epsilon \sim p(\epsilon)}\left[\nabla_\phi \log p_\theta(y \mid g(\phi, \epsilon, y))\right] \end{gathered}\] Now we can use Monte Carlo Estimation. By sampling $\epsilon$ from $p(\epsilon)$, which is $\mathcal{N}(0,I)$:
\[\begin{gathered} \mathbb{E}_{\epsilon \sim \mathcal{N}(0, I)}\left[\nabla_\phi \log p_\theta(y \mid g(\phi, \epsilon, y))\right] \\ \approx \frac{1}{L} \sum_{l=1}^L \nabla_\phi \log p_\theta\left(y \mid g\left(\phi, \epsilon^{(l)}, y\right)\right) \end{gathered}\] where $\epsilon^{(1)}, \ldots, \epsilon^{(L)} \sim \mathcal{N}(0, I)$
Why Monte Carlo works now?
Recall our building
\[\mathbb{E}_{x \sim q_\phi(x \mid y)}\left[\log p_\theta(y \mid x)\right]=\mathbb{E}_{\epsilon \sim p(\epsilon)}\left[\log p_\theta(y \mid g(\phi, \epsilon, y))\right]\] Sample $\epsilon\sim p(\epsilon)$ to estimate $\left[\log p_\theta(y \mid g(\phi, \epsilon, y))\right]$ is the same as sample $x \sim q_\phi(x \mid y)$ to estimate $\left[\log p_\theta(y \mid x)\right]$. Initially, $q_\phi\left(x \mid y\right)$ might output random mean, samples from $q_\phi$ don’t reconstruct well, causing high loss. But during the training, $q_\phi\left(x \mid y\right)$ concentrates around $x$ values where $p_\theta\left(y \mid x\right)$ is large, reaching our final goal $q_\phi(x \mid y) \approx p_\theta(x \mid y)$. This is actually related to importance sampling.
For simplicity, we set $L=1$.
\[\begin{gathered} \nabla_\phi \log p_\theta(y \mid g(\phi, \epsilon, y))\\ =\nabla_\phi \log p_\theta(y \mid \mu_\phi(y)+\sigma_\phi(y) \cdot \epsilon) \end{gathered}\] where $\epsilon \sim \mathcal{N}(0, I)$ is sampled once per gradient computation.
Now our update trajectory in expectation item is
\[\nabla_\phi \log p_\theta(y \mid \mu_\phi(y)+\sigma_\phi(y) \cdot \epsilon)\] We can build $\mu_\phi$ and $\sigma_\phi$ as neural networks, take this item as loss function, sample $\epsilon \sim \mathcal{N}(0, I)$, given a fixed $\theta$, this equation is solvable.
But wait, $\theta$ also a unknown variable.
True, we need to optimize both $\theta$ and $\phi$ simultaneously, recall our objective:
\[\max _{\theta, \phi} \operatorname{ELBO}(\theta, \phi ; y)\] We can make joint gradient descent . In each gradient step:
- Update $\phi: \phi^{(t+1)}=\phi^{(t)}+\alpha_\phi \cdot \nabla_\phi$ ELBO
- Update $\theta: \theta^{(t+1)}=\theta^{(t)}+\alpha_\theta \cdot \nabla_\theta$ ELBO
Thus we can learn $\phi$ and $\theta$ simultaneously.
Note: In this case two parameters cooperate (both improve the same goal), it looks like EM algorithm but actually not. An adversarial case is GAN where two parameters against each other.
]]>